MTS Helpdesk
Question:
When it comes to validation, analytical test methods used for medicinal products based on biological molecules can be a bit ‘tricky’ to deal with. Coming up with a suitable design for the validation protocol can be quite difficult. In particular, the choice of what parameters to investigate, and the design of the associated experiments. What are the main reasons for this?
Answer:
One of the main features that means biomolecules are more difficult is their size and associated complex structure. They are generally big molecules when compared to more traditional pharmaceutical active ingredients and this means that they not only contain more functional groups but also the orientation of the parts of the molecule in space is more complicated.
Size Matters
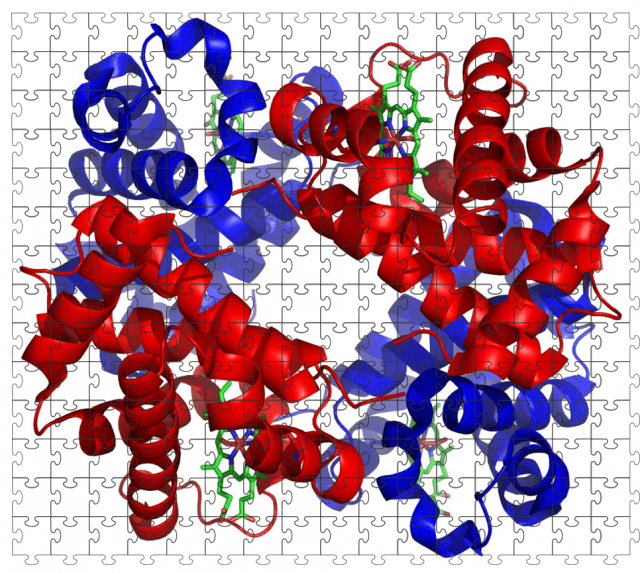
In the fairly typical, small molecule pharmaceutical, aspirin, there are only 9 carbon atoms present, with a total mass of just 180 Daltons. Contrast this with an example of a very large biopharmaceutical, a monoclonal antibody. Monoclonal antibodies, such as bevacizumab, have many more carbon atoms than the little aspirin molecule at over 6000 and have a total mass of approximately 149 kilodaltons, over 800 times bigger than aspirin. Of course not all biopharmaceuticals are as big as monoclonal antibodies but they still often have large and complex structures.
Jigsaw Puzzle Analogy
What this means when it comes to testing methods is that if we wanted to test aspirin tablets, then it is conceivable that a single test method, such as a chromatographic separation, could be used to provide all the information we require. Using a jigsaw puzzle analogy, the test method could provide different pieces of the puzzle that gives us the information we want, such as the amount of the aspirin molecule present in the tablets, and the amounts of other undesirable impurity molecules, to result in a full picture of what is in the sample. However, if we look at a biopharmaceutical sample, our puzzle pieces will be much smaller.
A single test method will not be enough to provide all the information that we want because there is so much more to consider. The puzzle pieces from one test method may build up an important picture of a particular attribute, perhaps the biological activity, or the glycans present, but we will need multiple test methods to really describe the biomolecule in terms of attributes such as its higher order structure, variants and impurities present.

Validation
The aim of validation is to demonstrate, typically via experiments, that the method is suitable for its intended use. The reason that ‘bio’ test methods are tricky to validate is that they are complex methods. Method validation for biopharmaceuticals usually follows the ICH guidance Q2(R1) which lists a number of parameters which should be investigated for different types of methods namely: Accuracy; Precision; Specificity; Detection limit; Quantitation limit; Linearity; Range and Robustness. These are called ‘validation characteristics’ in the ICH guidance but are now more commonly referred to as ‘method performance characteristics’ since they don’t just relate to validation activities.
If we have a look at this list we can pick out some of the most troublesome characteristics for ‘bio’ test methods. The quantitative parameters are often the most challenging. Accuracy, in particular, may be very difficult.
Accuracy
A common problem is that there is often no reference material available that is suitable for the test and therefore it is not possible to compare the outcome of any experiments to a true value. The purpose of the accuracy investigation is to show that to show that the value determined by the test is close to the true value, often shown as how close an arrow is to the centre of a target. If you don’t know the true value, then you can’t assert that the test method is accurate.
Examples would be: analysis of variants where it is impossible to isolate samples of each variant in order to check exactly how much is there; or impurities methods, where you may not definitively know what the impurity is, never mind have a sample of it to provide a true value. This forces us to make assumptions about the biomolecule and what the test is actually measuring such as assuming that the area of a peak in an electropheragram or a chromatogram actually represents how much is there compared to the other peaks.
It is not actually possible to investigate accuracy so we use other strategies to build up as much evidence as possible to show that the method is probably okay.
Quantitation Limit
Another characteristic affected by the lack of a true value is the quantitation limit. Without reference material we have to use dilutions to try to establish the most likely lowest level at which the method generates reasonable results.
Linearity
Linearity is also problematic. Many ‘bio’ test methods are not actually linear, the most common example being a biological assay which typically has a sigmoidal relationship. So really what we want to look at is the suitability of the calibration technique in the method. This is usually very complex for a bioassay but even for a more straightforward physiochemical method which does have a linear relationship it can still be challenging. To create this sort of plot where we compare the amount of the material in question with the response obtained by following the method, we will need a range of concentrations.
If we want to validate a method for a variant or an impurity, as we discussed previously, we are very unlikely to have reference material which would to allow changing the concentration of this component only and keeping the rest of the sample consistent over the full range required. What we will probably do instead is do perform dilutions on a sample of our material to achieve the range. This is not representative of the method, nor really is injecting different amounts instead of changing the concentration.
Another challenge of the calibration method is when area percent normalisation is used for quantification. In the example of an electropherogram from a capillary electrophoresis method shown below, the area of all the peaks would be assigned as 100% and each individual peak would then be calculated as a percentage of the total. This assumes a linear relationship right down to the origin of the calibration where the response and concentration both equal zero. It is difficult to prove this assumption through experiments when reference material is not available.
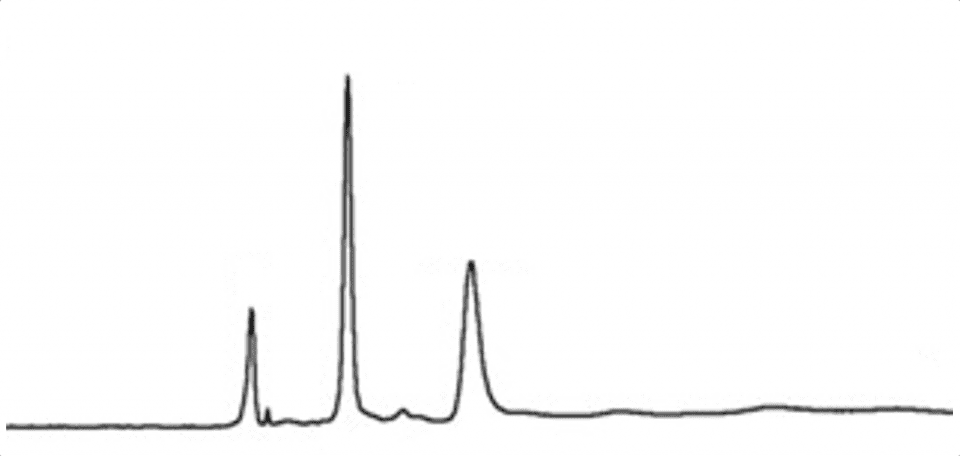

We have just looked at a few of the challenges when validating ‘bio’ methods to give a taste of the problems and underlying reasons.
What’s the takeaway message?
#1: Size matters, biomolecules are generally larger than traditional drug molecules and thus their structure is more complex,
#2: The structural complexity of biomolecules means that a range of methods is required to test them, no one method can provide all the required information, and
#3: Validation of ‘bio’ test methods is tricky because the methods often necessitate assumptions that are hard to prove experimentally.
If you want to validate one of these methods it will be easier if you have a very good understanding of how the method works, including all its assumptions and limitations.
Submit a Question to the MTS Helpdesk
Do you have a question for our expert relating to the topic of analytical chemistry? Use our enquiry form to send it to us.
Related content from our Resources Library
Services related to this content that we offer
Method validation for biomolecules is covered fully in our course, ‘Validation, Verification and Transfer of Methods for Biopharmaceutical Analysis‘.
Visit our courses page for a full list of available courses and the schedule of currently available course dates.